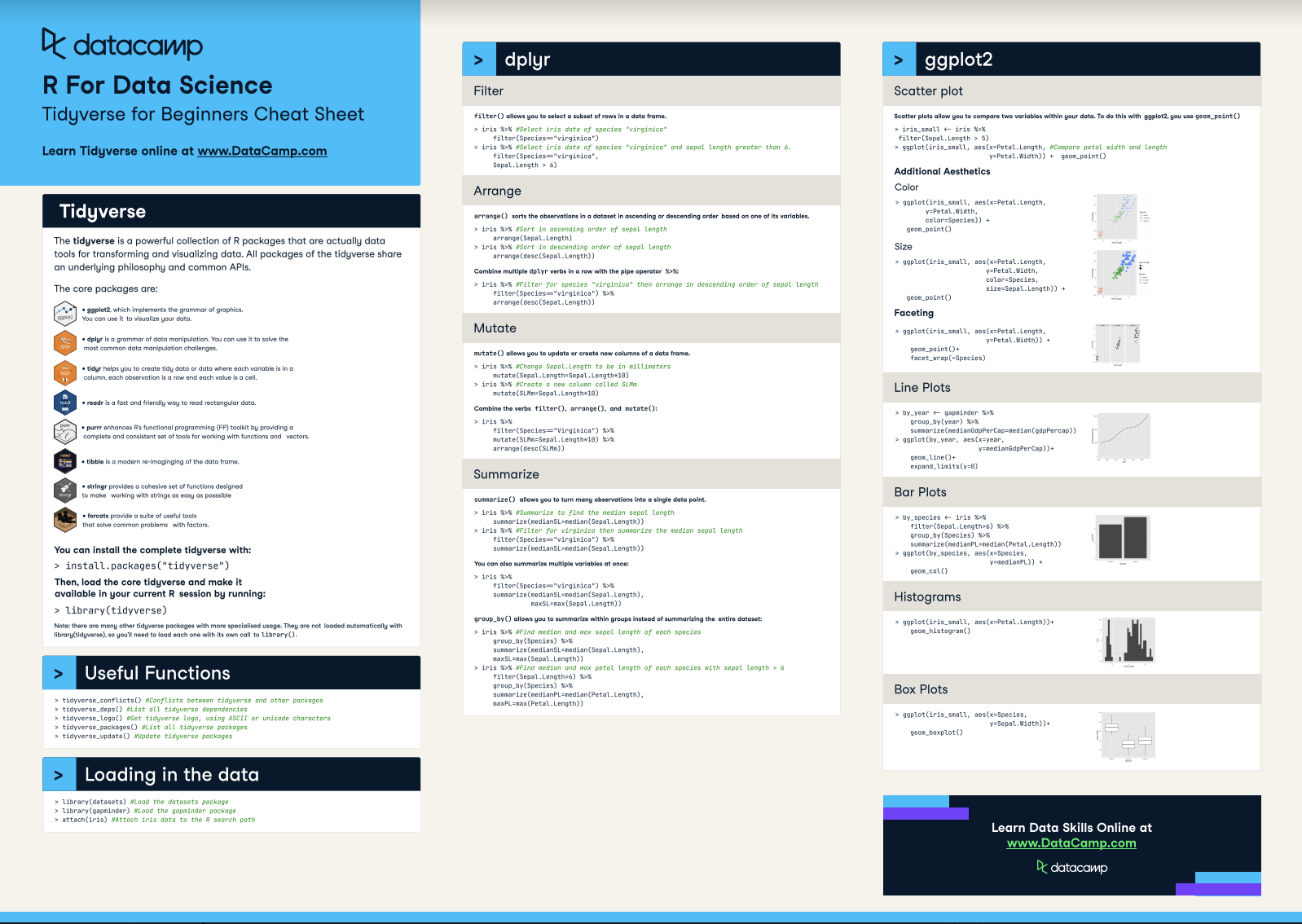
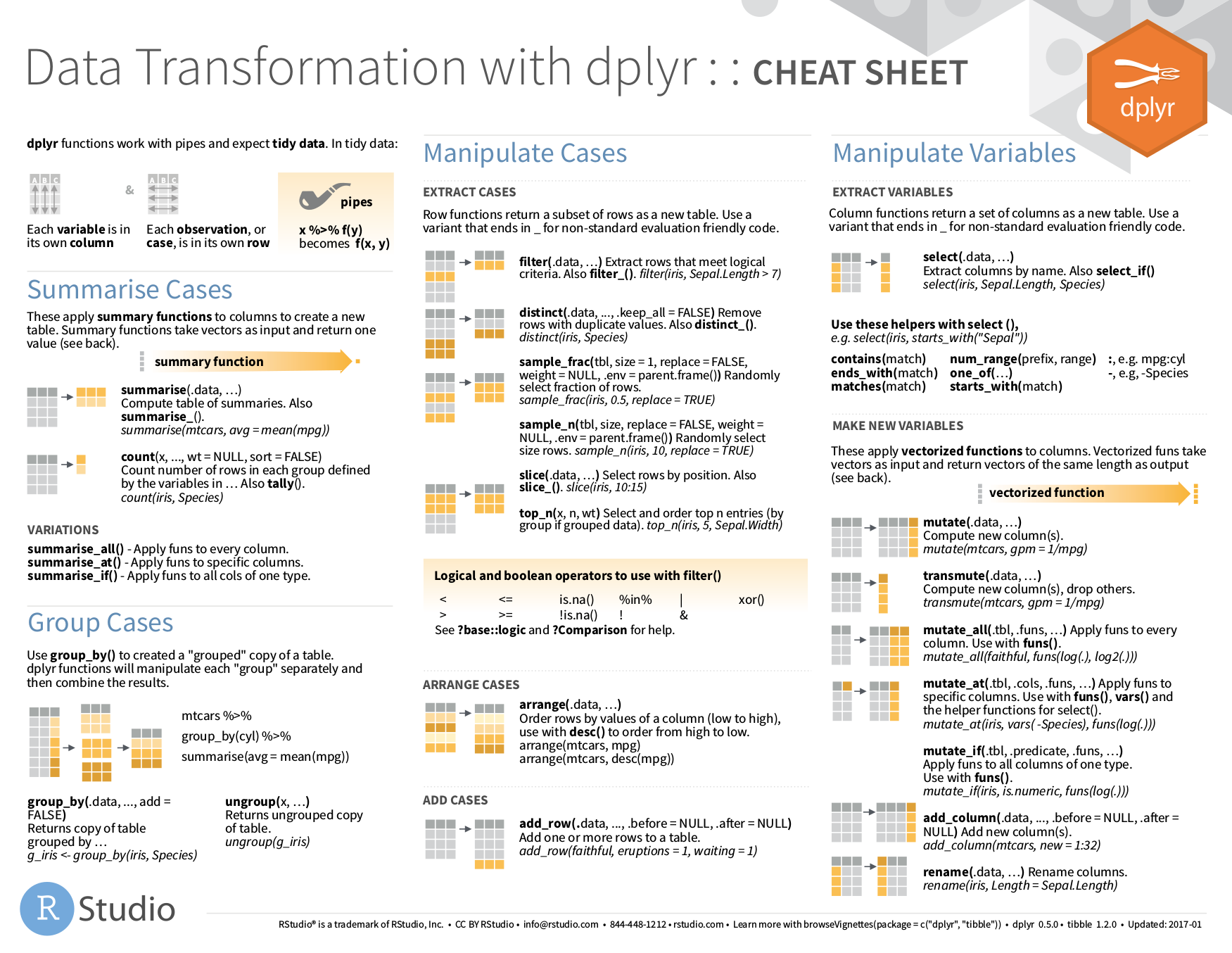
  
97.3 # The _if() variants apply a predicate function (a function that # returns TRUE or FALSE) to determine the relevant subset of # columns. 97.3 # -> starwars %>% summarise ( across ( height : mass, ~ mean (. 97.3 # You can also supply selection helpers to _at() functions but you have # to quote them with vars(): starwars %>% summarise_at ( vars ( height : mass ), mean, na.rm = TRUE ) #> # A tibble: 1 × 2 #> height mass #> #> 1 174. 97.3 # -> starwars %>% summarise ( across ( c ( "height", "mass" ), ~ mean (. # The _at() variants directly support strings: starwars %>% summarise_at ( c ( "height", "mass" ), mean, na.rm = TRUE ) #> # A tibble: 1 × 2 #> height mass #> #> 1 174. Name collisions in the new columns are disambiguated using a unique suffix. vars is named, a new column by that name will be created. Similarly, vars() accepts named and unnamed arguments. If a function is unnamed and the name cannot be derived automatically, funs argument can be a named or unnamed list. The names of the functions are used to name the new columns Ĭoncatenating the names of the input variables and the names of theįunctions, separated with an underscore "_". vars is of the form vars(a_single_column)) and. The names of the input variables are used to name the new columns įor _at functions, if there is only one unnamed variable (i.e., If there is only one unnamed function (i.e. Input variables and the names of the functions. Summary(df3) # we use summary() for many many other purposesĬlasses 'data.table' and 'ame': 20 obs. Comparing the outputs of read.csv(x) and fread(x) refers to the parent directory of your current directory, which is the /home directory. If your current directory is /home/desktop, then. can be used to refer to the parent directory. 
in the file path simply refers to the current working directory, so it can be dropped. It’s super intelligent and fast (reads gigabytes of data in just a few seconds). I always use fread() from the data.table to read data now. R has a lot of built-in datasets type data() in the console to see what dataests are available. Try typing sleep in your console and ?sleep for more info on this dataset. The sleep dataset is actually a built-in dataset in R. # READ: assign the output return by read.csv("data/sleep.csv") into df1ĭf2 <- fread("./data/sleep.csv") # fread() from library(data.table)ĭf3 <- fread("data/sleep.csv") # same as above # same as df1 <- read.csv("data/sleep.csv") Use library() to load packages at the top of each R script.ĭf1 <- read.csv("./data/sleep.csv") # base R read.csv() function It takes me many hours to research, learn, and put together tutorials. Consider being a patron and supporting my work?ĭonate and become a patron: If you find value in what I do and have learned something from my site, please consider becoming a patron. Get source code for this RMarkdown script here. Creating new variables/columns and reassigning in data.tables with :=.Compute summary statistics and apply functions to j by groups.Supercharging your workflow with data.table().Compute summary statistics with summarize() or summarise().Sorting or arranging data rows with arrange().Create new columns/variables with mutate().Filtering or subsetting data/rows with filter().Manipulating datasets with dplyr (a package in tidyverse).Writing/saving dataframes or datatables as csv files.Using $ and to extract elements using their names.Comparing the outputs of read.csv(x) and fread(x).Consider being a patron and supporting my work?. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
